Presslee et al. 2019
used mitochondrial DNA to assess relationships of tree sloths and their extinct relatives. “Results from phylogenetic analysis of these datasets differ substantially from morphology-based concepts.”
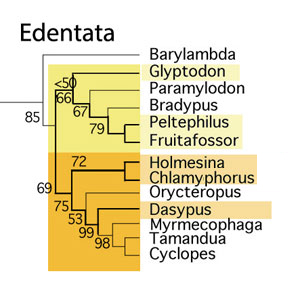
Figure 2. Subset of the LRT focusing on the Edentata. Armored taxa are color tinted and their branches are thicker.
No, kidding!
The Presslee team essentially flipped the clade Edentata upside-down and twisted it around, nesting the small arboreal and ant-loving taxa at the base and the basal herbivorous giants as derived. This conclusion is based on morphological testing in the large reptile tree (LRT, 1500 taxa; subset Fig. 1)
‘Palaeoproteomics’
is the term used to describe the applications of proteomics ( the analysis of sets of proteins) to ancient materials.
The validated outgroup taxon in the LRT, Barylambda,
was not tested by Presslee’s team.
The invalid outgroup they chose, Dasypus (the armadillo)
is a derived taxon in the LRT.
Summary
This study was a waste of time and paper. Phylogenetic analysis yields false positives when invalid outgroup taxa are employed. The software does not know any better than to follow instructions. Colleagues, please don’t trust DNA. Don’t choose an outgroup taxon. Let a wider gamut study that includes 1500 candidates choose a validated one for you.
References
Presslee S et al. (19 co-authors) 2019. Palaeoproteomics resolves sloth relationships. Nature Ecology & Evolution Online access.
